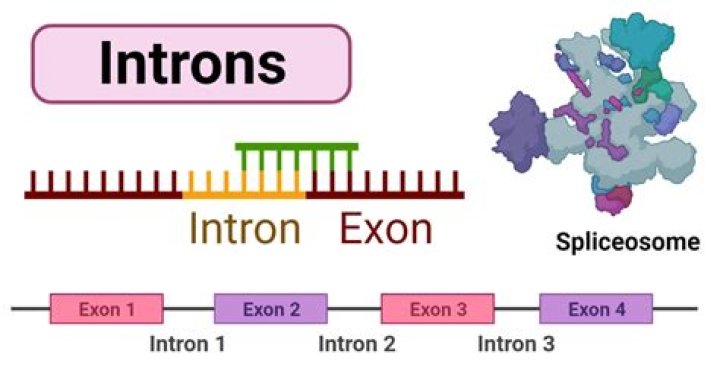
Why are introns cut out

Olivia Owen
Published Mar 19, 2026
Not only do the introns not carry information to build a protein, they actually have to be removed in order for the mRNA to encode a protein with the right sequence. If the spliceosome fails to remove an intron, an mRNA with extra “junk” in it will be made, and a wrong protein will get produced during translation.
Why is important to cut out the introns?
Introns create extra work for the cell because they replicate with each division, and cells must remove introns to make the final messenger RNA (mRNA) product. Organisms have to devote energy to get rid of them.
What is the purpose of introns?
Introns, from this perspective, have a profound purpose. They serve as hot spots for recombination in the formation of new combinations of exons. In other words, they are in our genes because they have been used during evolution as a faster pathway to assemble new genes.
What happens if an intron is not removed?
During the process of splicing, introns are removed from the pre-mRNA by the spliceosome and exons are spliced back together. If the introns are not removed, the RNA would be translated into a nonfunctional protein. Splicing occurs in the nucleus before the RNA migrates to the cytoplasm.Do introns have to be removed?
It is vital for the introns to be removed precisely, as any left-over intron nucleotides, or deletion of exon nucleotides, may result in a faulty protein being produced. This is because the amino acids that make up proteins are joined together based on codons, which consist of three nucleotides.
Why are introns important to evolution?
Evolutionary advantages of introns include the possibility to create new genes by cutting and pasting exons from existing genes or to diversify the protein output of a single gene by splicing the exons together in different ways.
Why is it important to remove the introns before proceeding with translation?
Not only do the introns not carry information to build a protein, they actually have to be removed in order for the mRNA to encode a protein with the right sequence. If the spliceosome fails to remove an intron, an mRNA with extra “junk” in it will be made, and a wrong protein will get produced during translation.
Are introns junk?
Introns are ubiquitous in eukaryotic transcripts. They are often viewed as junk RNA but the huge energetic burden of transcribing, removing, and degrading them suggests a significant evolutionary advantage. Ostensibly, an intron functions within the host pre-mRNA to regulate its splicing, transport, and degradation.Where do introns get removed?
Introns are removed from primary transcripts by cleavage at conserved sequences called splice sites. These sites are found at the 5′ and 3′ ends of introns. Most commonly, the RNA sequence that is removed begins with the dinucleotide GU at its 5′ end, and ends with AG at its 3′ end.
Are intron mutations bad?Mutations in these sequences may lead to retention of large segments of intronic DNA by the mRNA, or to entire exons being spliced out of the mRNA. These changes could result in production of a nonfunctional protein. An intron is separated from its exon by means of the splice site.
Article first time published onHow do introns enhance gene expression?
Introns can increase transcript levels by affecting the rate of transcription, nuclear export, and transcript stability. Moreover, introns can also increase the efficiency of mRNA translation. … Rather, each intron-gene combination might undergo its own unique mixture of processes that lead to the perceptible outcome.
Why do we have introns and exons?
The mixing and matching of exons from the same gene can lead to proteins with different functions. … Therefore, introns are a way to generate different proteins or different amounts of proteins that are unique to a cell type. Introns might also allow for faster evolution.
Why are there no introns in prokaryotes?
Prokaryotic cells need and use virtually all of there DNA to synthesize protein and RNA molecules. After DNA is transcribed into RNA in a eukaryotic cell, it’s further edited in the nucleus, part of the RNA is cut out, this part cut out is the introns. It’s absent in prokaryotes because there’s no nucleus.
What happens to the introns?
After transcription of a eukaryotic pre-mRNA, its introns are removed by the spliceosome, joining exons for translation. … Other intron products have long half-lives and can be exported to the cytoplasm, suggesting that they have roles in translation.
Where do introns go after splicing?
Where do the introns go after splicing in eukaryotes? – Quora. Usually, they don’t go anywhere. They get broken down into their component nucleotides by nuclear exonucleases (that almost sounded poetic). In humans, the bulk of this activity is carried out by XRN2.
What happens to the cut out intron after alternative RNA splicing?
It is a sequence that codes for the binding of RNA polymerase to the DNA. … What happens to the cut-out intron after alternative RNA splicing? A. It is added back to the mRNA.
What is the role of retained introns?
We describe the roles of introns that are retained in mature mRNAs to regulate normal cellular development in plants and animals. … This process involves removal of introns between adjacent exons by a massive RNA- protein complex known as the spliceosome [35], comprising five RNAs and over 200 proteins [36, 37].
Which enzyme removes introns?
Spliceozymes: Ribozymes that Remove Introns from Pre-mRNAs in Trans.
How do introns evolve?
The introns-late concept held that introns emerged only in eukaryotes and new introns have been accumulating continuously throughout eukaryotic evolution. … Conversely, splicing errors gave rise to alternative splicing, a major contribution to the biological complexity of multicellular eukaryotes.
Why are introns not junk DNA?
Introns are cut, or ‘spliced,’ out of the mRNA before it gets translated into a protein. In other words, they aren’t used to make the final protein product. At first introns might look like junk, but lots of them aren’t. … But only 3% or so of all DNA has the information to make proteins.
What happens to the cut out introns?
The pre-mRNA molecule thus goes through a modification process in the nucleus called splicing during which the noncoding introns are cut out and only the coding exons remain. … Splicing produces a mature messenger RNA molecule that is then translated into a protein.
What would happen if introns were not removed during RNA processing?
If introns were not edited out of the RNA strand, the RNA strand would probably have many problems. Errors would most likely occur in the instruction code for amino acids and proteins and the cell therefore would not get the amount of proteins needed. … A site where RNA polymerase can bind to begin transcription.
Are introns always the same?
For example, introns are extremely common within the nuclear genome of jawed vertebrates (e.g. humans and mice), where protein-coding genes almost always contain multiple introns, while introns are rare within the nuclear genes of some eukaryotic microorganisms, for example baker’s/brewer’s yeast (Saccharomyces …
How much of our DNA is introns?
Only about 1 percent of DNA is made up of protein-coding genes; the other 99 percent is noncoding.
Why are introns longer than exons?
Recent studies report that alternatively spliced exons tend to occur in longer introns, which is attributed to the length constraints for splice site pairing for the two major splicing mechanisms, intron definition versus exon definition.
What are introns vs exons?
The parts of the gene sequence that are expressed in the protein are called exons, because they are expressed, while the parts of the gene sequence that are not expressed in the protein are called introns, because they come in between the exons.
Do mutations occur in introns?
Here we review evidence from mRNA analysis and entire genomic sequencing indicating that pathogenic mutations can occur deep within the introns of over 75 disease-associated genes.
Do mutations in introns affect phenotype?
In addition to pathological mutations sensu stricto, introns also harbour functional polymorphisms that can influence the expression of the genes that host them. Some of these intronic variants may also confer susceptibility to disease or otherwise modulate the genotype-phenotype relationship.
What effect would a mutation in an intron have on the expression of gene?
Introns and Human Health The intron sequences that affect mRNA accumulation are redundant and dispersed, so a point mutation or even a large deletion in the intron would probably not significantly reduce the expression of the gene unless splicing was disrupted.
Is there an advantage for Intronless genes?
Intronless genes can serve as beacons in analyses of gene function and evolution. For example, intronless genes as a model, compared with intron-containing homologs, can enable an inverse approach to studying the considerable roles of introns, which are only found in eukaryotes [1].
What are indels in regards to DNA sequences?
Indel is a molecular biology term for an insertion or deletion of bases in the genome of an organism. … An indel inserts and deletes nucleotides from a sequence, while a point mutation is a form of substitution that replaces one of the nucleotides without changing the overall number in the DNA.



